Publications & Research
Welcome to my Publications & Research page! Here, I highlight my authored and collaborative research contributions.
You can also explore my work on Web of Science and Google Scholar.
My primary focus is on leveraging advanced image analysis tools to extract quantifiable features from pathology images, aiming to discover prognostically relevant patterns directly from routine pathology slides. This includes studying molecular alterations and the tumor immune microenvironment.
For a short summary, please click on a project title.
Research Summary
1. During Postdoc (MD Anderson Cancer Center)
Conducted Research
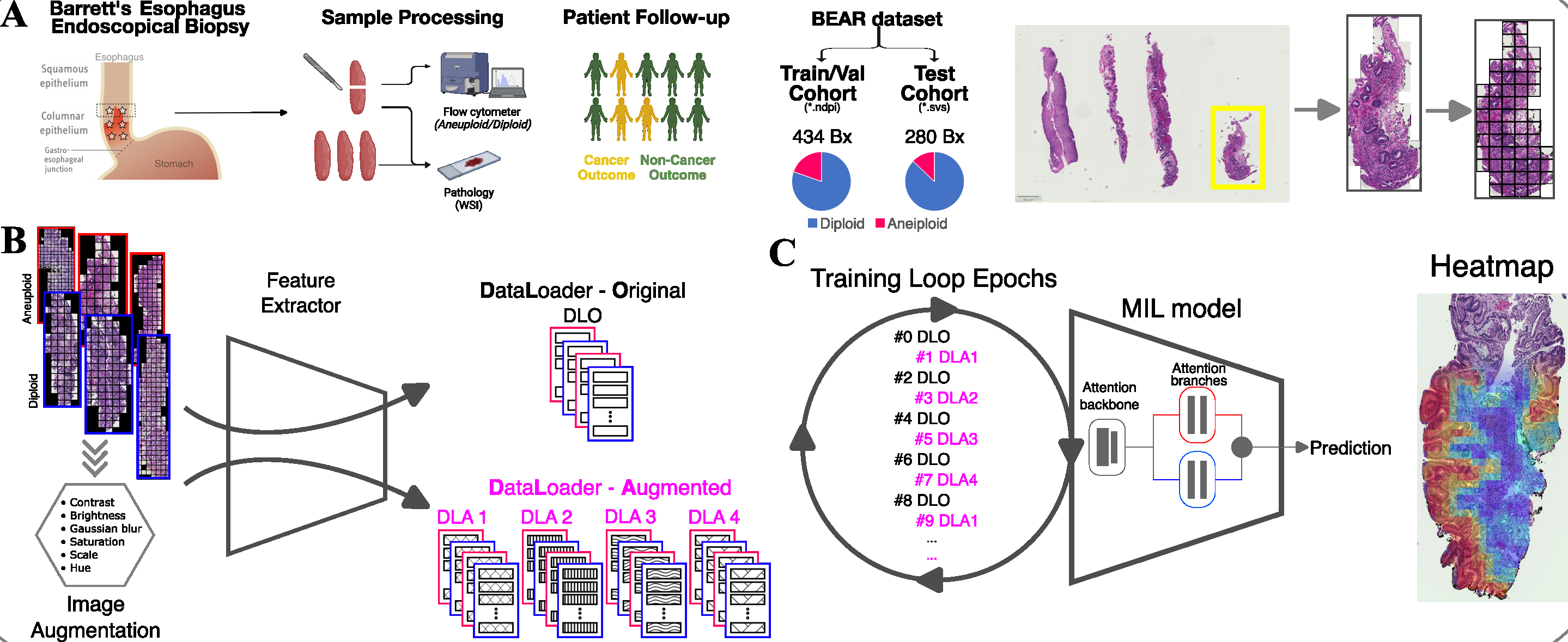
Here, I present the research projects I led as the first author.During my postdoctoral fellowship in the United States, I have been conducting research in computational pathology. My main project, "Deep learning-based whole-slide image analysis for immune ecology and cancer progression prediction in Barrett’s esophagus," aims to: 1- Predict aneuploidy directly from endoscopic biopsy images. 2- Investigate the spatial distribution of immune responses to uncover tumor-immune interactions. 3- Identify features relevant to cancer progression. 4- Develop a prognostic prediction tool. This fellowship was funded by the funded by SNF in postdoc.Mobility grant schema. While this project is ongoing, key milestones have been presented at scientific conferences. For more details, visit this page.
Collaborations & Contributions
Besides my main project, I have been involved in multiple projects in my lab which I contribute with my expertise in pathology, morphology, immunemicroenvironment or pathologenesis, and also utilisation of softwares, algorithms in computarinal pathology research. THe collaborations which I involved and reached a milestone that could be published are listed below with brief desriptions.- Spatial Transcriptomics Deconvolution: Developed an algorithm to deconvolve spatial transcriptomics data, submitted to AACR 2025. - DeMixT 2.0: A Deconvolution Framework for Sparse Sequencing Data Using Embedded Negative Binomial Distribution.
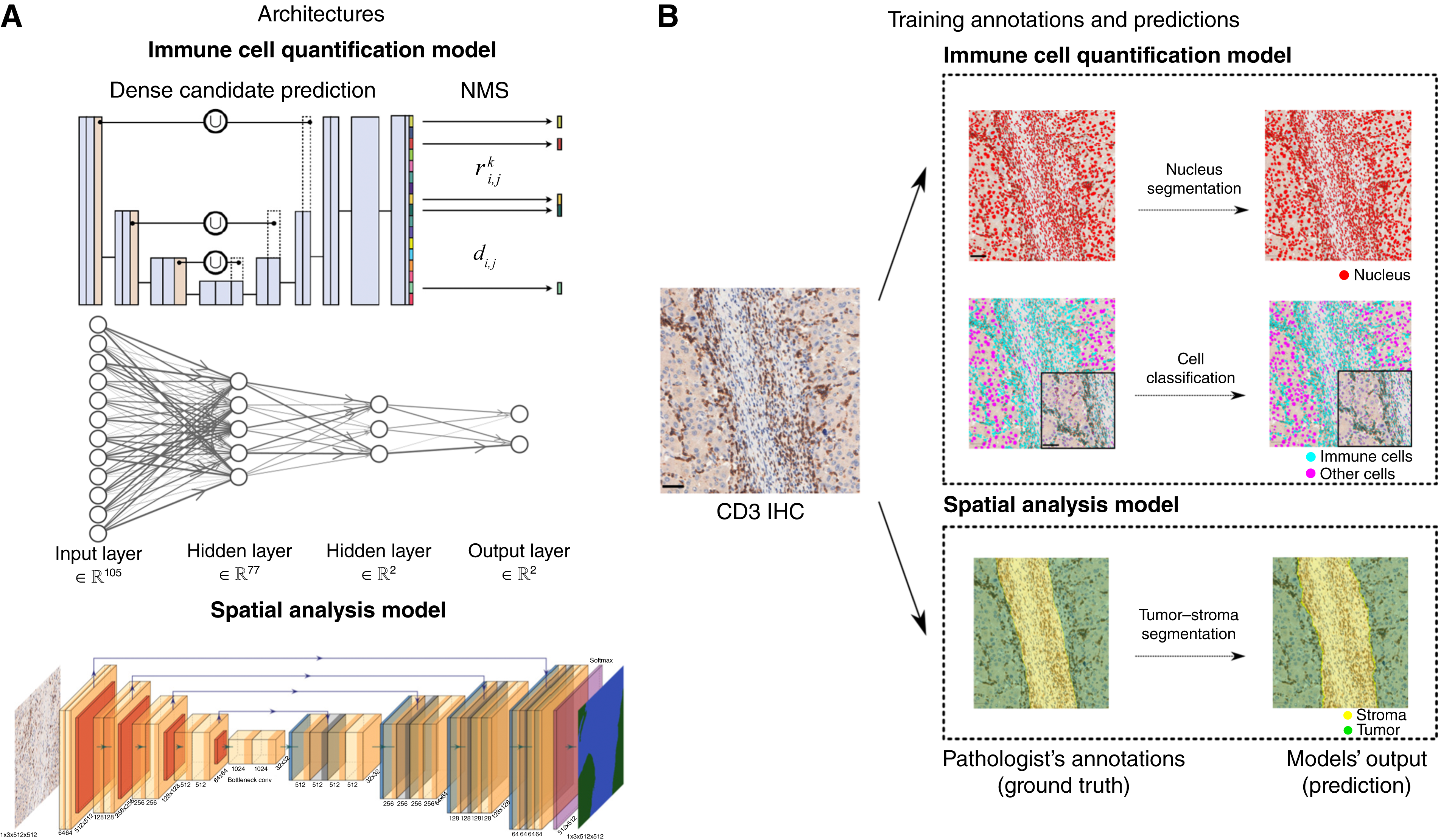
- Tumor Immune Microenvironment Segmentation: Developed a tool to annotate spatial transcriptomics spots for lung cancer, submitted to AACR 2025. - TMEseg: Tumor Microenvironment Segmentation from Histology Images.
- Breast Cancer Response Prediction: Built an AI-based tool to predict response to treatment in invasive breast carcinoma, accepted to SABCS 2024. - AI-Derived Tumor-Infiltrating Lymphocytes Enhance Prediction of Pathologic Complete Response in Early-Stage Triple-Negative Breast Cancer.
- Ductal Carcinoma Progression Prediction: Created a tool for progression prediction in ductal carcinoma in situ, submitted to AACR 2025. - AI-Derived Tumor-Infiltrating Lymphocytes Predict Invasive Breast Cancer Recurrence.
- Morphological Feature Analysis: Revealed cancer progression features in ductal carcinoma in situ, accepted to npj Precision Oncology. - A Morphometric Signature to Identify Ductal Carcinoma In Situ with Low Risk of Progression.
- Tumor Heterogeneity Sampling Tool: Developed a computational pathology tool for reproducible sampling of heterogeneous tumor regions, accepted to USCAP 2025. - AI-STORM: AI-Guided Sampling of Tissues for Optimal Representative Morphology.